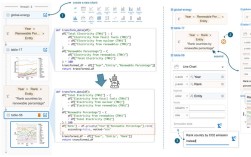
Of course! Here is a comprehensive guide to creating dendrograms in Python, covering the most popular libraries, their use cases, and a complete workflow.
What is a Dendrogram?
A dendrogram is a tree-like diagram that visualizes the arrangement of clusters produced by hierarchical clustering. It shows how each cluster is composed by drawing links between clusters based on their similarity.
The key to reading a dendrogram is the y-axis (distance):
- The vertical lines represent clusters.
- The height of a fusion (where two vertical lines meet) represents the distance (or dissimilarity) between the two clusters being joined. A higher fusion means the clusters were more dissimilar and joined later in the process.
- The horizontal lines just link clusters, and their length has no special meaning.
Method 1: The Easiest Way with scipy and matplotlib
This is the most common and straightforward method for creating a dendrogram from a pre-computed distance matrix or data. It's perfect for understanding the clustering structure.
Key Libraries:
scipy.cluster.hierarchy: Provides the core functions for hierarchical clustering (linkage) and dendrogram plotting (dendrogram).scipy.spatial.distance.pdist: Computes the pairwise distances between observations in a dataset.matplotlib.pyplot: Used for displaying the plot.
Step-by-Step Example:
Let's create a dendrogram for some sample data.
import numpy as np
import matplotlib.pyplot as plt
from scipy.cluster.hierarchy import dendrogram, linkage
from scipy.spatial.distance import pdist
# 1. Generate some sample data
# Let's create 3 distinct groups of points
np.random.seed(42)
group1 = np.random.randn(10, 2) + [2, 2]
group2 = np.random.randn(10, 2) + [-2, -2]
group3 = np.random.randn(10, 2) + [2, -2]
# Combine them into a single array
X = np.vstack((group1, group2, group3))
# 2. Compute the linkage matrix
# 'pdist' calculates the pairwise distances.
# 'linkage' performs the hierarchical clustering using a method (e.g., 'ward', 'single', 'complete')
# 'ward' is a popular choice that minimizes the variance within clusters.
distance_matrix = pdist(X, metric='euclidean')
Z = linkage(distance_matrix, method='ward')
# 3. Plot the dendrogram
plt.figure(figsize=(10, 7))"Dendrogram for Sample Data")
plt.xlabel("Sample index")
plt.ylabel("Distance (Ward)")
# The 'dendrogram' function can truncate the display for large datasets
dendrogram(
Z,
truncate_mode='lastp', # Show only the last p merged clusters
p=12, # Show the last 12 merged clusters
show_leaf_counts=True, # Show the number of points in each cluster
leaf_rotation=90., # Rotate leaf labels
leaf_font_size=12.,
show_contracted=True # To get a distribution impression in truncated branches
)
plt.show()
Explanation of Key linkage Methods:
The choice of method in linkage drastically changes the shape of the dendrogram.
'ward': Minimizes the variance of the clusters being merged. It's a good, default choice.'single': Uses the minimum distance between any two points in the two clusters. Can lead to "chaining" (long, stringy clusters).'complete': Uses the maximum distance between any two points in the two clusters. Tends to produce compact, spherical clusters.'average': Uses the average of all pairwise distances.
Method 2: The Integrated Way with scikit-learn and scipy
scikit-learn is the go-to library for machine learning. While its AgglomerativeClustering class is excellent for performing hierarchical clustering and getting flat cluster labels, it doesn't have a built-in dendrogram plotter. The standard practice is to use scikit-learn for the clustering logic and scipy for the visualization.
This approach is useful when you are already working within the scikit-learn ecosystem.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.cluster import AgglomerativeClustering
from scipy.cluster.hierarchy import dendrogram, linkage
# 1. Generate sample data using scikit-learn
X, y_true = make_blobs(n_samples=50, centers=3, cluster_std=1.0, random_state=42)
# 2. Perform hierarchical clustering with scikit-learn
# Note: scikit-learn's AgglomerativeClustering does not return a linkage matrix
# directly, but we can get the children_ attribute.
# However, for plotting, it's often easier to just re-compute the linkage with scipy.
# Let's do it the "easy" way by recomputing the linkage matrix.
Z = linkage(X, method='ward', metric='euclidean')
# 3. Plot the dendrogram
plt.figure(figsize=(12, 6))"Dendrogram from scikit-learn data")
dendrogram(Z, truncate_mode='lastp', p=10, show_leaf_counts=True)
plt.xlabel("Sample index")
plt.ylabel("Distance (Ward)")
plt.show()
# --- Bonus: How to get flat clusters from the dendrogram ---
# You can decide on a distance threshold or a number of clusters (k)
# and cut the dendrogram to get flat clusters.
# Option A: Cut by distance threshold
distance_threshold = 10
clusters = fcluster(Z, t=distance_threshold, criterion='distance')
print(f"Number of clusters for distance threshold {distance_threshold}: {len(np.unique(clusters))}")
# Option B: Cut by number of clusters (k)
k = 3
clusters_k = fcluster(Z, t=k, criterion='maxclust')
print(f"Cluster assignments for k={k}: {clusters_k}")
# Visualize the flat clusters
plt.figure(figsize=(8, 6))
plt.scatter(X[:, 0], X[:, 1], c=clusters_k, cmap='viridis', s=50, alpha=0.8)f"Flat Clusters (k={k})")
plt.xlabel("Feature 1")
plt.ylabel("Feature 2")
plt.show()
Method 3: The "Publication-Ready" Way with plotly
For interactive and visually appealing dendrograms, especially for web-based applications or presentations, plotly is an excellent choice. It allows you to hover over points to see more information.
First, you'll need to install it:
pip install plotly
import numpy as np
import plotly.figure_factory as ff
from scipy.cluster.hierarchy import linkage, fcluster
# 1. Generate sample data
np.random.seed(42)
group1 = np.random.randn(20, 2) + [2, 2]
group2 = np.random.randn(20, 2) + [-2, -2]
group3 = np.random.randn(20, 2) + [2, -2]
X = np.vstack((group1, group2, group3))
# 2. Compute the linkage matrix
Z = linkage(X, method='ward', metric='euclidean')
# 3. Get cluster labels for coloring the dendrogram
# Let's say we want to show 3 main clusters
labels = fcluster(Z, t=3, criterion='maxclust')
# 4. Create the Plotly dendrogram
fig = ff.create_dendrogram(
X,
linkagefun=lambda x: linkage(x, method='ward', metric='euclidean'),
color_threshold=10, # Distance at which to color the clusters
labels=[f'Point {i}' for i in range(X.shape[0])] # Optional: custom labels
)
# Update layout for better appearance
fig.update_layout(
width=800,
height=500,text="Interactive Dendrogram with Plotly",
xaxis_title="Sample Index",
yaxis_title="Distance (Ward)"
)
# Show the plot
fig.show()
Summary: Which Method to Use?
| Method | Best For | Pros | Cons |
|---|---|---|---|
scipy + matplotlib |
Quick and simple visualization. Standard for data analysis and exploration. | - Easy to use. - No extra dependencies beyond the scientific Python stack. - Highly customizable with matplotlib. |
- Static plots. - Can be slow with very large datasets. |
scikit-learn + scipy |
Machine learning workflows. When you need to perform clustering and get labels for other ML tasks. | - Integrates seamlessly with other scikit-learn tools (e.g., pipelines, GridSearchCV).- Easy to get flat cluster labels ( fcluster). |
- Requires using two libraries for a full workflow. - scikit-learn doesn't natively plot dendrograms. |
plotly |
Interactive dashboards and presentations. When you need to explore data by hovering and zooming. | - Fully interactive (zoom, pan, hover). - Modern, publication-quality aesthetics. - Great for web-based applications. |
- Requires an extra library installation. - Can be overkill for simple, static plots. |