Of course! Here is a complete guide to creating a Mandelbrot set in Python, starting with a simple version and moving to a more optimized and visually appealing one using the numpy and matplotlib libraries.
What is the Mandelbrot Set?
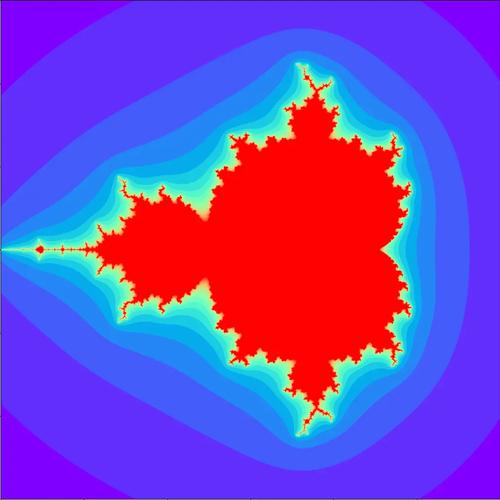
The Mandelbrot set is a famous fractal, a shape that exhibits self-similarity at all scales. It's defined as the set of complex numbers c for which the following sequence does not diverge to infinity:
- Start with
z₀ = 0. - Iterate the formula:
z_{n+1} = z_n² + c. - If the magnitude of
z(i.e.,|z|) remains bounded (less than 2) after many iterations, the numbercis in the Mandelbrot set.
To visualize it, we map a region of the complex plane (e.g., from -2 to 1 on the real axis and -1 to 1 on the imaginary axis) to a grid of pixels. For each pixel, we treat its coordinates as a complex number c and run the iteration test. The number of iterations it takes for |z| to exceed 2 determines the pixel's color.
Method 1: The Simple, Pure Python Approach
This version is great for understanding the core logic. It's easy to read but can be slow for high-resolution images.
The Code
import matplotlib.pyplot as plt
def mandelbrot_simple(width, height, max_iterations):
"""
Generates the Mandelbrot set using a simple, pure Python approach.
"""
# Create a 2D array to store the number of iterations for each pixel
iterations_grid = [[0 for _ in range(width)] for _ in range(height)]
# Define the bounds of the complex plane we want to visualize
# The Mandelbrot set is roughly centered at (-0.5, 0) in the complex plane
x_min, x_max = -2.0, 1.0
y_min, y_max = -1.0, 1.0
for row in range(height):
for col in range(width):
# Map the pixel coordinates to a complex number c
x0 = x_min + (col / width) * (x_max - x_min)
y0 = y_min + (row / height) * (y_max - y_min)
c = complex(x0, y0)
# Iterate the Mandelbrot formula
z = 0j
iteration = 0
# We check if |z| is less than 2, which is equivalent to |z|^2 < 4
while (z.real * z.real + z.imag * z.imag) <= 4 and iteration < max_iterations:
z = z * z + c
iteration += 1
iterations_grid[row][col] = iteration
return iterations_grid
# --- Main execution ---
if __name__ == "__main__":
IMG_WIDTH = 800
IMG_HEIGHT = 600
MAX_ITERATIONS = 100
print("Calculating Mandelbrot set...")
grid = mandelbrot_simple(IMG_WIDTH, IMG_HEIGHT, MAX_ITERATIONS)
# Create a plot
plt.figure(figsize=(10, 7.5))
plt.imshow(grid, extent=[-2.0, 1.0, -1.0, 1.0], cmap='hot', origin='lower')
plt.colorbar(label='Iterations until divergence')
plt.title("Mandelbrot Set (Simple Python)")
plt.xlabel("Real axis")
plt.ylabel("Imaginary axis")
plt.show()
How to Run It
- Make sure you have
matplotlibinstalled:pip install matplotlib - Save the code as a Python file (e.g.,
mandelbrot_simple.py). - Run it from your terminal:
python mandelbrot_simple.py
Explanation
-
mandelbrot_simplefunction: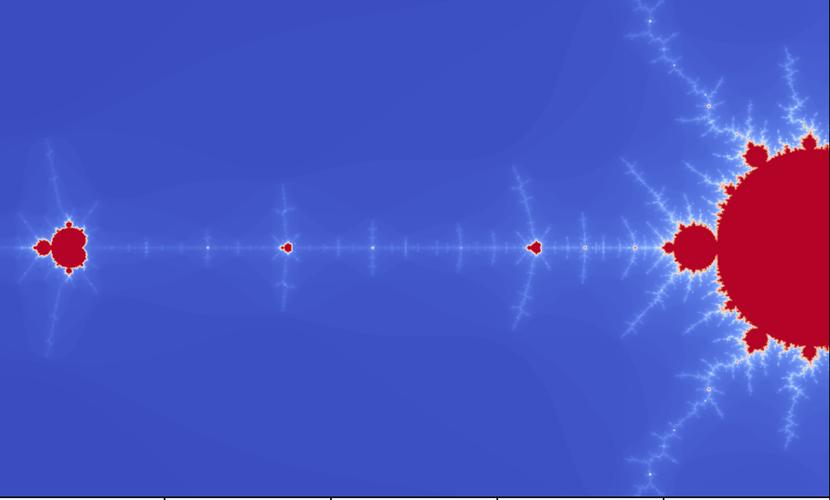 (图片来源网络,侵删)
(图片来源网络,侵删)- It takes
width,height, andmax_iterationsas input. - It initializes a 2D list
iterations_gridto store the result for each pixel. - It loops through every pixel
(row, col). - Mapping to Complex Plane: It maps the pixel's integer coordinates to a floating-point coordinate
(x0, y0)within our desired bounds (x_mintox_max,y_mintoy_max). This(x0, y0)becomes our complex numberc. - Iteration Loop: It initializes
z = 0and then repeatedly applies the formulaz = z*z + c. It stops if|z|exceeds 2 or if it reachesmax_iterations. - Store Result: The final iteration count is stored in
iterations_grid.
- It takes
-
Main Block:
- Sets the desired image dimensions and maximum iterations.
- Calls the function to generate the data.
- Plotting:
matplotlib.pyplot.imshow()is used to create the image. It takes our 2D list and assigns colors based on the values.cmap='hot'gives a classic "hot" color map (black -> red -> yellow -> white).extenttellsimshowthe real-world coordinates of the image.
Method 2: The Optimized NumPy Approach
This version is significantly faster because it uses NumPy's vectorized operations to perform calculations on the entire grid at once, avoiding slow Python loops.
The Code
import numpy as np
import matplotlib.pyplot as plt
def mandelbrot_numpy(width, height, max_iterations):
"""
Generates the Mandelbrot set using NumPy for fast vectorized operations.
"""
# Create a 2D array of complex numbers representing the c values for each pixel
x_min, x_max = -2.0, 1.0
y_min, y_max = -1.0, 1.0
x = np.linspace(x_min, x_max, width)
y = np.linspace(y_min, y_max, height)
# Create a meshgrid, which is a grid of coordinate points
X, Y = np.meshgrid(x, y)
# Combine the real and imaginary parts into a single complex array C
C = X + 1j * Y
# Create a copy of C to use as Z (the initial value of Z is 0)
Z = np.zeros_like(C)
# Create an array to store the iteration counts
iterations = np.zeros(C.shape, dtype=int)
# Create a mask for points that haven't diverged yet
# We start by assuming no points have diverged
not_diverged = np.ones(C.shape, dtype=bool)
for i in range(max_iterations):
# Only update Z for the points that haven't diverged yet
Z[not_diverged] = Z[not_diverged]**2 + C[not_diverged]
# Check for divergence: |Z|^2 > 4
# The new mask is the intersection of the old mask and the non-diverged condition
not_diverged = not_diverged & (np.abs(Z) <= 4)
# Increment the iteration count for points that are still in the set
iterations[not_diverged] += 1
return iterations
# --- Main execution ---
if __name__ == "__main__":
IMG_WIDTH = 800
IMG_HEIGHT = 600
MAX_ITERATIONS = 100
print("Calculating Mandelbrot set with NumPy...")
grid = mandelbrot_numpy(IMG_WIDTH, IMG_HEIGHT, MAX_ITERATIONS)
# Create a plot
plt.figure(figsize=(10, 7.5))
plt.imshow(grid, extent=[-2.0, 1.0, -1.0, 1.0], cmap='hot', origin='lower')
plt.colorbar(label='Iterations until divergence')
plt.title("Mandelbrot Set (NumPy)")
plt.xlabel("Real axis")
plt.ylabel("Imaginary axis")
plt.show()
Explanation of Differences
- Vectorization: Instead of looping through pixels, we create a grid of complex numbers
Cusingnp.linspaceandnp.meshgrid. This single arrayCholds all thecvalues for our image. - Boolean Masking: The key to the optimization is the
not_divergedboolean array. This array acts as a mask. In each iteration, we only perform calculations (Z = Z**2 + C) on the elements ofZandCwherenot_divergedisTrue. - Efficiency: NumPy performs these array operations in highly optimized, pre-compiled C code, which is orders of magnitude faster than a Python
forloop. This makes the NumPy version capable of handling much higher resolutions and more iterations in a reasonable time.
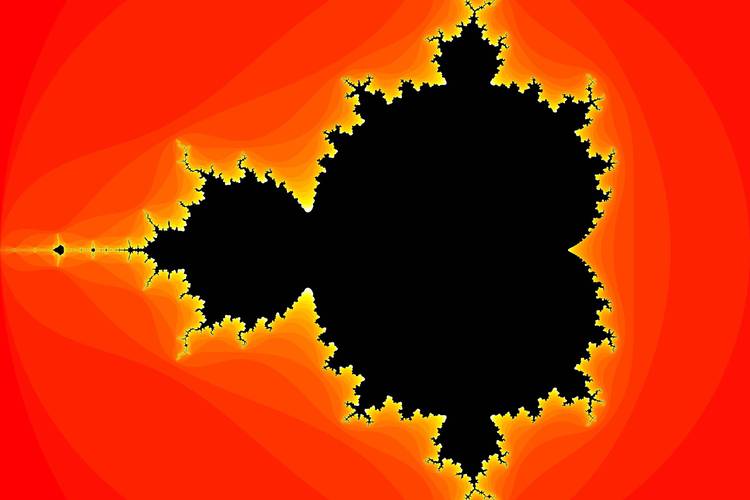
Going Further: Adding Color and Zooming
The hot colormap is simple. For more beautiful images, we can create a custom colormap that maps the iteration count to a smooth color gradient.
Smooth Coloring
The simple method gives a "banding" effect. We can use a continuous coloring algorithm for a smoother look. A common technique is to use the fractional part of the iteration count.

import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import LinearSegmentedColormap
def mandelbrot_smooth(width, height, max_iterations):
# ... (same setup as the NumPy version) ...
x_min, x_max = -2.0, 1.0
y_min, y_max = -1.0, 1.0
x = np.linspace(x_min, x_max, width)
y = np.linspace(y_min, y_max, height)
X, Y = np.meshgrid(x, y)
C = X + 1j * Y
Z = np.zeros_like(C)
iterations = np.zeros(C.shape, dtype=int)
not_diverged = np.ones(C.shape, dtype=bool)
for i in range(max_iterations):
Z[not_diverged] = Z[not_diverged]**2 + C[not_diverged]
# Check for divergence: |Z|^2 > 4
diverged_now = (np.abs(Z) > 4) & not_diverged
iterations[diverged_now] = i + 1 # Points that diverged in this step
not_diverged = not_diverged & (np.abs(Z) <= 4)
# --- Smooth coloring ---
# For points that did not diverge, set their iteration count to max_iterations
iterations[not_diverged] = max_iterations
# Calculate the smoothed value
# The formula is: nu = n + 1 - log(log(|z_n|)) / log(2)
# We add a small epsilon to avoid log(0)
log_zn = np.log(np.abs(Z) + 1e-10)
nu = np.log(log_zn / np.log(2)) / np.log(2)
smooth_iterations = iterations + 1 - nu
# Cap the values at max_iterations
smooth_iterations[iterations == max_iterations] = max_iterations
return smooth_iterations
# --- Custom Colormap ---
colors = [(0, 0, 0.5), (0, 0, 1), (0, 0.5, 1), (0, 1, 1), (0.5, 1, 0.5), (1, 1, 0), (1, 0.5, 0), (1, 0, 0), (0.5, 0, 0)]
cmap_name = 'mandelbrot_colormap'
cm = LinearSegmentedColormap.from_list(cmap_name, colors, N=256)
# --- Main execution ---
if __name__ == "__main__":
IMG_WIDTH = 1200
IMG_HEIGHT = 800
MAX_ITERATIONS = 256 # Higher iterations for smoother coloring
print("Calculating smooth Mandelbrot set...")
grid = mandelbrot_smooth(IMG_WIDTH, IMG_HEIGHT, MAX_ITERATIONS)
plt.figure(figsize=(12, 9))
plt.imshow(grid, extent=[-2.0, 1.0, -1.0, 1.0], cmap=cm, origin='lower', interpolation='bilinear')
plt.colorbar(label='Smoothed Iterations')
plt.title("Smooth Mandelbrot Set with Custom Colormap")
plt.xlabel("Real axis")
plt.ylabel("Imaginary axis")
plt.show()
Zooming In
To zoom in, you just need to change the x_min, x_max, y_min, and y_max values. For example, to zoom into the "seahorse valley":
# In the main execution block, replace the extent and x/y min/max with: x_min, x_max = -0.75, -0.74 y_min, y_max = 0.1, 0.11 # And adjust the plot extent accordingly: # plt.imshow(..., extent=[x_min, x_max, y_min, y_max], ...)
This will create a new image focused on that specific region of the fractal, revealing infinite detail.