Of course! Here is a detailed explanation and a complete, runnable Python implementation of the Metropolis-Hastings algorithm, a foundational Markov Chain Monte Carlo (MCMC) method.
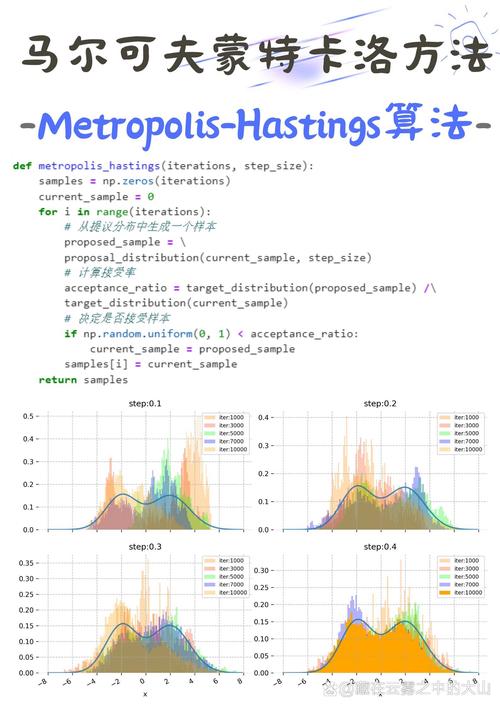
What is the Metropolis Algorithm?
The Metropolis algorithm is a method used to generate a sequence of random samples from a probability distribution for which direct sampling is difficult. This is incredibly common in fields like physics, statistics, and machine learning.
Core Idea: Imagine you want to understand the shape of a complex, multi-dimensional mountain range (your target probability distribution), but you can't just survey the whole thing. Instead, you can take a random walk. At each step, you propose a new direction. If the new spot is higher (more probable), you definitely move there. If it's lower (less probable), you might still move there, but with a probability that depends on how much lower it is. Over time, the places you visit will be distributed according to the "height" of the mountain range.
Key Concepts
-
Target Distribution (
p(x)): This is the probability distribution we want to sample from. Often, we only know it up to a normalizing constant. That is, we knowp(x) = f(x) / Z, wheref(x)is a function we can evaluate, butZ(the integral off(x)over all space) is impossible to compute. The Metropolis algorithm cleverly bypasses the need forZ. -
Proposal Distribution (
q(x' | x)): This distribution suggests a new statex'given the current statex. A common and simple choice is a symmetric distribution, like a Gaussian centered at the current point:x' = x + ε, whereε ~ Normal(0, σ²)For this symmetric proposal,q(x' | x) = q(x | x'). (图片来源网络,侵删)
(图片来源网络,侵删) -
Acceptance Ratio (
A): This is the heart of the algorithm. It decides whether to accept or reject the proposed movex'.A = min( 1, p(x') / p(x) )Because ourp(x)is often unnormalized (f(x)), we can use the ratio of the unnormalized densities, which conveniently cancels out the unknownZ:A = min( 1, f(x') / f(x) ) -
Markov Chain: The sequence of samples generated forms a Markov chain, meaning the next state only depends on the current state, not the entire history.
The Algorithm Steps
Let's say we want to sample from a target distribution p(x).
- Initialize: Choose a starting point
x₀and sett = 0. - Iterate: For each step
tfrom 1 toN(the total number of samples): a. Propose: Generate a new candidate samplex'from the proposal distributionq(x' | x_t). (e.g.,x' = x_t + random.normal(0, step_size)). b. Calculate Acceptance Ratio: ComputeA = min(1, p(x') / p(x_t)). c. Accept or Reject:- Generate a uniform random number
ufrom[0, 1]. - If
u <= A, accept the new sample: setx_{t+1} = x'. - Else, reject the new sample: set
x_{t+1} = x_t(the chain stays in its current place).
- Generate a uniform random number
- Collect Samples: After running the chain for
Nsteps, you have a sequence of samples{x₀, x₁, ..., x_N}.
Important Considerations
- Burn-in: The initial part of the chain (e.g., the first 1000-5000 steps) is often influenced by the arbitrary starting point
x₀and may not be representative of the target distribution. This period is called the "burn-in" period and is usually discarded. - Thinning/Correlation: Consecutive samples in the chain are correlated. To get a more independent set of samples, you can "thin" the chain by only keeping every
k-th sample (e.g., every 10th sample). - Step Size (): The standard deviation of the proposal Gaussian is crucial. If it's too small, the chain will take tiny steps and explore the space very slowly (high correlation). If it's too large, most proposals will be in low-probability regions and will be rejected, also leading to slow exploration. The goal is to achieve an "acceptance rate" of around 20-50%.
Python Implementation: Sampling from a 2D Gaussian
Let's use the Metropolis algorithm to sample from a 2D Gaussian (bivariate normal) distribution with a known mean and covariance. This is a great example because we can compare our MCMC samples to the true distribution.
We will use numpy for numerical operations and matplotlib for plotting.
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# --- 1. Define the Target Distribution ---
# We'll sample from a 2D Gaussian distribution.
# The "unnormalized" probability density function (PDF) is all we need.
# We don't need the normalization constant (1 / (2*pi*sigma_x*sigma_y*sqrt(1-rho^2)))
def target_distribution(x, y):
"""
Unnormalized PDF of a 2D Gaussian distribution.
"""
# Parameters of our target distribution
mean = np.array([0.0, 0.0])
cov = np.array([[1.0, 0.8], [0.8, 1.0]]) # High correlation
# Vectorized calculation
pos = np.array([x, y])
# The exponent part of the Gaussian PDF
exponent = -0.5 * (pos - mean).T @ np.linalg.inv(cov) @ (pos - mean)
# We return the exponential part. The constant multiplier is not needed.
return np.exp(exponent)
# --- 2. The Metropolis-Hastings Algorithm ---
def metropolis_sampler(num_samples, start_pos, step_size):
"""
Runs the Metropolis-Hastings algorithm to sample from a target distribution.
Args:
num_samples (int): The total number of samples to generate.
start_pos (np.array): The starting position [x, y].
step_size (float): The standard deviation for the Gaussian proposal.
Returns:
np.array: A (num_samples, 2) array of the generated samples.
"""
# Initialize the chain with the starting position
current_pos = np.array(start_pos, dtype=float)
samples = [current_pos]
# Pre-calculate the log of the target at the current position to avoid repeated calculations
log_target_current = np.log(target_distribution(*current_pos))
for i in range(num_samples):
# a. Propose a new position
# The proposal is symmetric: q(x'|x) = N(x, step_size^2)
proposal = current_pos + np.random.normal(0, step_size, size=2)
# b. Calculate the acceptance ratio (in log space for numerical stability)
log_target_proposal = np.log(target_distribution(*proposal))
# For a symmetric proposal, log(q(x'|x)) - log(q(x|x')) = 0
# So, log(A) = log(min(1, p(x')/p(x))) = min(0, log(p(x')) - log(p(x)))
log_acceptance_ratio = log_target_proposal - log_target_current
# c. Accept or reject
# Generate a uniform random number
u = np.log(np.random.uniform(0, 1))
if u < log_acceptance_ratio:
# Accept the proposal
current_pos = proposal
log_target_current = log_target_proposal # Update the log target for the new position
# Add the current position to the samples (whether we moved or not)
samples.append(current_pos)
return np.array(samples)
# --- 3. Run the Sampler and Visualize Results ---
# Parameters for the MCMC
NUM_SAMPLES = 20000
START_POS = [-5.0, -5.0] # Start from a point far from the mean
STEP_SIZE = 1.5 # Tune this parameter for good acceptance rate (~20-50%)
print("Running Metropolis sampler...")
samples = metropolis_sampler(NUM_SAMPLES, START_POS, STEP_SIZE)
print("Sampler finished.")
# --- 4. Post-processing and Visualization ---
# a. Discard burn-in samples (e.g., first 20%)
burn_in = int(0.2 * len(samples))
processed_samples = samples[burn_in:]
# b. Plot the results
plt.style.use('seaborn-v0_8-whitegrid')
fig, ax = plt.subplots(figsize=(10, 8))
# Plot the MCMC samples
ax.plot(processed_samples[:, 0], processed_samples[:, 1], 'o', markersize=1, alpha=0.5, label='MCMC Samples')
# Plot the true distribution as a contour plot for comparison
x = np.linspace(-4, 4, 200)
y = np.linspace(-4, 4, 200)
X, Y = np.meshgrid(x, y)
Z = target_distribution(X, Y)
ax.contour(X, Y, Z, levels=15, colors='k', alpha=0.3, linestyles='dashed')
# Mark the starting position
ax.plot(START_POS[0], START_POS[1], 'go', markersize=10, label='Start Position')
# Mark the mean of the target distribution
ax.plot(0, 0, 'r*', markersize=15, label='True Mean')
ax.set_title(f'Metropolis-Hastings Sampling Results (N={NUM_SAMPLES}, Step Size={STEP_SIZE})')
ax.set_xlabel('X-axis')
ax.set_ylabel('Y-axis')
ax.legend()
ax.axis('equal')
plt.show()
# --- 5. Diagnostics ---
# Let's check the acceptance rate
accepted_moves = 0
for i in range(1, len(samples)):
if not np.array_equal(samples[i], samples[i-1]):
accepted_moves += 1
acceptance_rate = accepted_moves / (len(samples) - 1)
print(f"\nDiagnostics:")
print(f"Acceptance Rate: {acceptance_rate:.2%}")
# Plot the trace (the value of the chain over time) for the first dimension
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(12, 8), sharex=True)
# Trace plot
ax1.plot(samples[:, 0], alpha=0.5)
ax1.set_title('Trace Plot (X-dimension)')
ax1.set_ylabel('X value')
ax1.axvline(x=burn_in, color='r', linestyle='--', label='End of Burn-in')
ax1.legend()
# Autocorrelation plot
from statsmodels.tsa.stattools import acf
lags = 100
autocorr = acf(samples[burn_in:, 0], nlags=lags, fft=True)
ax2.plot(autocorr)
ax2.set_title('Autocorrelation Plot (X-dimension)')
ax2.set_xlabel('Lag')
ax2.set_ylabel('Autocorrelation')
ax2.axhline(y=0, color='k', linestyle='-', alpha=0.3)
ax2.axhline(y=0.2, color='r', linestyle='--', alpha=0.5, label='Threshold')
ax2.legend()
plt.tight_layout()
plt.show()
How to Interpret the Output
-
Main Scatter Plot:
- You will see a cloud of points that densely fills the shape of the underlying 2D Gaussian.
- The dashed contour lines match the shape of your MCMC samples, showing that the algorithm successfully learned the target distribution.
- The "Start Position" is far away, and you can visually trace how the chain "walks" from that corner towards the high-probability region near the mean (the red star).
-
Diagnostics:
- Acceptance Rate: A rate between 20% and 50% is generally a good sign that your
step_sizeis well-tuned. If it's too low, yourstep_sizeis too big. If it's too high (e.g., >90%), yourstep_sizeis too small. - Trace Plot: This shows the value of the
xcoordinate at each step of the chain. After the burn-in period (the red dashed line), the plot should look like "fat, fuzzy caterpillar"—randomly fluctuating around a stable mean. If it shows trends or slow drifts, the chain might not have converged yet. - Autocorrelation Plot: This shows how correlated a sample is with its past self. High autocorrelation means your samples are not very independent. The plot should drop to near zero relatively quickly. If it stays high for many lags, it indicates high correlation, and you might want to use "thinning" (keeping only every
k-th sample).
- Acceptance Rate: A rate between 20% and 50% is generally a good sign that your